| Orthocoronavirinae | |
|---|---|
| |
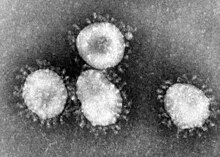
| Transmission electron micrograph of Avian coronavirus | |
| |
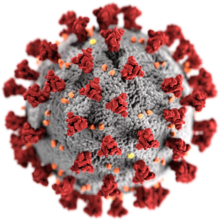
| Illustration of a SARS-CoV-2 virion[2] Red: spike proteins (S) Yellow: envelope proteins (E) Orange: membrane proteins (M)
| |
| Virus classification | |
| (unranked): | Virus |
| Realm: | incertae sedis |
| Kingdom: | incertae sedis |
| Phylum: | incertae sedis |
| Class: | incertae sedis |
| Order: | Nidovirales |
| Family: | Coronaviridae |
| Subfamily: | Orthocoronavirinae |
| Genera[1] | |
| |
| Synonyms[3][4] | |
| |
Coronaviruses are a group of RNA viruses.[5][6] They cause diseases in birds and mammals, including humans. These diseases can be mild, or they can be fatal. In humans and birds, they cause respiratory tract infections. Mild illnesses in humans include some cases of the common cold (which is also caused by other viruses, for example rhinoviruses). More lethal varieties can cause SARS, MERS, and COVID-19.
They are enveloped viruses with a positive-sense RNA genome.[7] The genome size of coronaviruses is about 26 to 32 kilobases,[8] which is extraordinarily large for an RNA virus. There are four major groups of coronaviruses, called alpha, beta, gamma, and delta.
The most famous coronavirus is one of the betas, the kind that causes Coronavirus disease 2019 (COVID-19) in humans.
The name "coronavirus" comes from the Latin word corona, meaning "crown" or "halo", and refers to how virions look under an electron microscopy (E.M.).[9] They have a fringe of large, bulbous surface projections looking like a crown. This morphology is created by the viral spike (S) peplomer, which are proteins on the surface of the virus. They decide which cells the virus can infect.
Proteins of coronaviruses are the spike (S), envelope (E), membrane (M), and nucleocapsid (N).
Diseases
[change | change source]Coronaviruses infect the upper respiratory and gastrointestinal tracts of mammals and birds. Six different strains of coronaviruses infect humans. These include:
- MERS-CoV
- SARS-CoV
- Severe acute respiratory syndrome coronavirus 2, which causes the disease coronavirus disease 2019 and is the cause of the COVID-19 pandemic (formerly referred as the "2019–20 coronavirus outbreak")
Coronaviruses are believed to cause many common colds in human adults. The significance and economic impact of coronaviruses is hard to assess. Unlike rhinoviruses (another common cold virus), human coronaviruses are easy to grow in the laboratory.
Structure
[change | change source]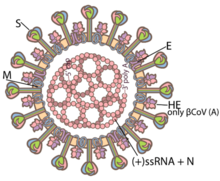
Coronaviruses are large, spherical particles with unique surface projections.[10] Their size is variable, averageing 80 to 120 nms.[11] The total molecular weight is on average 40,000 kDa. They are enclosed in an envelope studded with projecting protein molecules.[12] These layers protect the virus when it is outside the host cell.[13]
The viral envelope is made up of a lipid bilayer in which the membrane (M), envelope (E), and spike (S) structural proteins are anchored.[14] The ratio of E:S:M in the lipid bilayer is approximately 1:20:300.[15] The E and M protein are the structural proteins that combined with the lipid bilayer to shape the viral envelope and maintain its size. S proteins are needed for interaction with the host cells. But human coronavirus NL63 is peculiar in that its M protein has the binding site for the host cell, and not its S protein.[16] The diameter of the envelope is 85 nm.
History
[change | change source]The earliest reports of a coronavirus infection in animals occurred in the late 1920s, when an acute respiratory infection of domesticated chickens happened in North America.[17] Arthur Schalk and M.C. Hawn in 1931 made the first detailed report which described a new respiratory infection of chickens in North Dakota. It was not realized at the time that three related viruses were involved.[18]
Human coronaviruses were discovered in the 1960's using two different methods.[19][20][21] E.C. Kendall, Malcolm Bynoe, and David Tyrrel, working at the Common Cold Unit of the British Medical Research Council, collected a unique common cold virus B814 in 1961.[22][23][24] The virus could not be cultivated using the techniques which had successfully cultivated rhinoviruses, adenoviruses, and other known common cold viruses. Eventually, this was solved.[25] The new cultivating method was introduced to the lab by Bertil Hoorn.[26] The isolated virus, when put into the noses of volunteers, caused a cold. The virus was inactivated by ether which showed it had a lipid envelope.[22][27][28] The novel virus caused a cold in volunteers and, like B814, was inactivated by ether.[29]
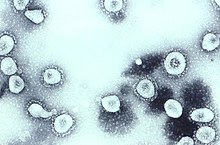
Scottish virologist June Almeida at St Thomas' Hospital, London, compared the structures of IBV, B814, and 229E in 1967.[30][31] Using transmission electron microscopy,the three viruses were shown to be similar in shape and have club-like spikes.[32] A research group at the National Institute of Health the same year isolated another member of this group of viruses.[33] Like B814, 229E, and IBV, the novel cold virus OC43 had distinctive club-like spikes when observed with the electron microscope.[34][35]
The IBV-like novel cold viruses looked like the mouse hepatitis virus. This new group of viruses were called "coronaviruses" from their crown-like appearance.[36][37] The coronavirus strain B814 was lost. It is not known which present human coronavirus it was.[38] Other human coronaviruses have since been identified, including SARS-CoV in 2003, HCoV NL63 in 2003, HCoV HKU1 in 2004, MERS-CoV in 2013, and SARS-CoV-2 in 2019.[39] There have been a large number of animal coronaviruses identified since the 1960s.
Related pages
[change | change source]References
[change | change source]- ↑ "Virus Taxonomy: 2018b Release". International Committee on Taxonomy of Viruses (ICTV). March 2019. Archived from the original on 4 March 2018. Retrieved 24 January 2020.
- ↑ Giaimo C (1 April 2020). "The Spiky Blob Seen Around the World". The New York Times. Archived from the original on 2 April 2020. Retrieved 6 April 2020.
- ↑ "2017.012-015S" (xlsx). International Committee on Taxonomy of Viruses (ICTV). October 2018. Archived from the original on 14 May 2019. Retrieved 24 January 2020.
- ↑ de Groot R.J. (2012). "Family Coronaviridae". In King A.M.Q. ' (ed.). Ninth Report of the International Committee on Taxonomy of Viruses. Elsevier, Oxford. pp. 806–828. ISBN 978-0-12-384684-6.
- ↑ International Committee on Taxonomy of Viruses. "ICTV Master Species List 2009 – v10" (xls).
- ↑ enveloped = has viral envelopes covering their protective protein capsids; positive sense = the RNA sequence may be directly translated into the desired viral proteins.
- ↑ One kilobase = 1000 base pairs
- ↑ virion = complete virus particle.
- ↑ Goldsmith CS, Tatti KM, Ksiazek TG, Rollin PE, Comer JA, Lee WW, et al. (February 2004). "Ultrastructural characterization of SARS coronavirus". Emerging Infectious Diseases. 10 (2): 320–6. doi:10.3201/eid1002.030913. PMC 3322934. PMID 15030705.
Virions acquired an envelope by budding into the cisternae and formed mostly spherical, sometimes pleomorphic, particles that averaged 78 nm in diameter (Figure 1A).
- ↑ Masters PS (2006). "The molecular biology of coronaviruses". Advances in Virus Research. 66: 193–292. doi:10.1016/S0065-3527(06)66005-3. ISBN 9780120398690. PMC 7112330. PMID 16877062.
- ↑ Lalchhandama K (2020). "The chronicles of coronaviruses: the electron microscope, the doughnut, and the spike". Science Vision. 20 (2): 78–92. doi:10.33493/scivis.20.02.03.
- ↑ Neuman BW, Kiss G, Kunding AH, Bhella D, Baksh MF, Connelly S, et al. (April 2011). "A structural analysis of M protein in coronavirus assembly and morphology". Journal of Structural Biology. 174 (1): 11–22. doi:10.1016/j.jsb.2010.11.021. PMC 4486061. PMID 21130884.
See Figure 10.
- ↑ Lai MM, Cavanagh D (1997). "The molecular biology of coronaviruses". Advances in Virus Research. 48: 1–100. doi:10.1016/S0065-3527(08)60286-9. ISBN 9780120398485. PMC 7130985. PMID 9233431.
- ↑ Cavanagh D, Mawditt K, Sharma M, Drury SE, Ainsworth HL, Britton P, Gough RE (August 2001). Schmidt A, Weber O, Wolff MH (eds.). "Detection of a coronavirus from turkey poults in Europe genetically related to infectious bronchitis virus of chickens". Avian Pathology. Birkhäuser Advances in Infectious Diseases BAID. 30 (4). Birkhäuser: 355–68. doi:10.1007/3-7643-7339-3_1. ISBN 978-3-7643-7339-9. PMC 7123520. PMID 19184921.
- ↑ Naskalska A, Dabrowska A, Szczepanski A, Milewska A, Jasik KP, Pyrc K (October 2019). "Membrane Protein of Human Coronavirus NL63 Is Responsible for Interaction with the Adhesion Receptor". Journal of Virology. 93 (19). doi:10.1128/JVI.00355-19. PMC 6744225. PMID 31315999.
- ↑ Estola T (1970). "Coronaviruses, a New Group of Animal RNA Viruses". Avian Diseases. 14 (2): 330–336. doi:10.2307/1588476. ISSN 0005-2086. JSTOR 1588476. PMID 4316767.
- ↑ "Il était une fois les coronavirus". Réalités Biomédicales (in French). 2020-03-27. Retrieved 2020-04-18.
- ↑ Kahn JS, McIntosh K (November 2005). "History and recent advances in coronavirus discovery". The Pediatric Infectious Disease Journal. 24 (11 Suppl): S223–7, discussion S226. doi:10.1097/01.inf.0000188166.17324.60. PMID 16378050.
- ↑ Mahase E (April 2020). "The BMJ in 1965". BMJ. 369: m1547. doi:10.1136/bmj.m1547. PMID 32299810.
- ↑ Monto AS (1984). "Coronaviruses". In Evans AS (ed.). Viral Infections of Humans. Springer US. pp. 151–165. doi:10.1007/978-1-4684-4727-9_7. ISBN 978-1-4684-4727-9.
((cite book)):|work=ignored (help) - ↑ 22.0 22.1 Kendall EJ, Bynoe ML, Tyrrell DA (July 1962). "Virus isolations from common colds occurring in a residential school". British Medical Journal. 2 (5297): 82–6. doi:10.1136/bmj.2.5297.82. PMC 1925312. PMID 14455113.
- ↑ Richmond C (2005-06-18). "David Tyrrell". BMJ : British Medical Journal. 330 (7505): 1451. doi:10.1136/bmj.330.7505.1451. PMC 558394.
- ↑ "Obituary Notices: Malcom Byone". British Medical Journal. 2 (5660): 827–829. 1969-06-28. doi:10.1136/bmj.2.5660.827. S2CID 220187042.
- ↑ Tyrrell DA, Bynoe ML (June 1965). "Cultivation of a novel type of common-cold virus in organ cultures". British Medical Journal. 1 (5448): 1467–70. doi:10.1136/bmj.1.5448.1467. PMC 2166670. PMID 14288084.
- ↑ Tyrrell DA, Fielder M (2002). Cold Wars: The Fight Against the Common Cold. Oxford University Press. pp. 93–95. ISBN 978-0-19-263285-2.
- ↑ Hagan WA, Bruner DW, Gillespie JH, Timoney JF, Scott FW, Barlough JE (1988). Hagan and Bruner's Microbiology and Infectious Diseases of Domestic Animals. Cornell University Press. p. 440. ISBN 978-0-8014-1896-9.
- ↑ Knapp, Alex. "The secret history of The first coronavirus". Forbes. Retrieved 2020-05-06.
- ↑ Hamre D, Procknow JJ (January 1966). "A new virus isolated from the human respiratory tract". Proceedings of the Society for Experimental Biology and Medicine. 121 (1): 190–3. doi:10.3181/00379727-121-30734. PMID 4285768. S2CID 1314901.
- ↑ "The woman who discovered the first coronavirus". Apr 14, 2020. Retrieved May 13, 2022 – via www.bbc.com.
- ↑ Almeida J (2008-06-26). "June Almeida (née Hart)". BMJ. 336 (7659): 1511.1–1511. doi:10.1136/bmj.a434. ISSN 0959-8138. PMC 2440895.
- ↑ Almeida JD, Tyrrell DA (April 1967). "The morphology of three previously uncharacterized human respiratory viruses that grow in organ culture". The Journal of General Virology. 1 (2): 175–8. doi:10.1099/0022-1317-1-2-175. PMID 4293939.
- ↑ McIntosh K, Becker WB, Chanock RM (December 1967). "Growth in suckling-mouse brain of "IBV-like" viruses from patients with upper respiratory tract disease". Proceedings of the National Academy of Sciences of the United States of America. 58 (6): 2268–73. Bibcode:1967PNAS...58.2268M. doi:10.1073/pnas.58.6.2268. PMC 223830. PMID 4298953.
- ↑ McIntosh K, Dees JH, Becker WB, Kapikian AZ, Chanock RM (April 1967). "Recovery in tracheal organ cultures of novel viruses from patients with respiratory disease". Proceedings of the National Academy of Sciences of the United States of America. 57 (4): 933–40. Bibcode:1967PNAS...57..933M. doi:10.1073/pnas.57.4.933. PMC 224637. PMID 5231356.
- ↑ Times, Harold M. Schmeck Jr Special To the New York (1967-05-05). "Six newly discovered viruses may explain cold: strains are similar to germ that causes a bronchial infection in chickens believed to be new group". The New York Times. ISSN 0362-4331. Retrieved 2020-04-25.
- ↑ Myint SH (1995). "Human Coronavirus Infections". In Siddell SG (ed.). The Coronaviridae. The Viruses. Springer US. pp. 389–401. doi:10.1007/978-1-4899-1531-3_18. ISBN 978-1-4899-1531-3. S2CID 80726096.
- ↑ Geller C, Varbanov M, Duval RE (November 2012). "Human coronaviruses: insights into environmental resistance and its influence on the development of new antiseptic strategies". Viruses. 4 (11): 3044–68. doi:10.3390/v4113044. PMC 3509683. PMID 23202515.
- ↑ Corman VM, Jores J, Meyer B, Younan M, Liljander A, Said MY, et al. (August 2014). "Coronaviruses". Viral Infections of Humans. Vol. 20. pp. 1319–22. doi:10.1007/978-1-4899-7448-8_10. ISBN 978-1-4899-7447-1. PMC 7122465. PMID 25075637.
The other OC strains and B814 that could not be adapted to mouse brain resisted adaptation to cell culture as well; these distinct viruses have since been lost and may actually have been rediscovered recently.
((cite book)):|journal=ignored (help) - ↑ Zhu N, Zhang D, Wang W, Li X, Yang B, Song J, et al. (February 2020). "A Novel Coronavirus from Patients with Pneumonia in China, 2019". The New England Journal of Medicine. 382 (8): 727–733. doi:10.1056/NEJMoa2001017. PMC 7092803. PMID 31978945.
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Viruses |
|
|---|---|
| Symptoms |
|
| Complications |
|
| Drugs |
|